Accountable: AI Must Come with Responsibility, Auditability, and Clear Ownership
When an AI system recommends that a doctor prescribe a specific antibiotic, and the doctor follows that advice without knowing why, who is responsible if the patient is harmed? When an industrial AI guides a manufacturing decision that ruins a $2M batch of steel, whose career is on the line? This is the accountability vacuum.
The Blame Game Nobody Wins
AI accountability fails in a predictable pattern. Developers blame the data. Users blame the black box. And somewhere in the middle, a real person pays the price.
The fundamental problem is that AI is probabilistic, not deterministic. Unlike traditional software, where you can trace a bug to a line of code, AI errors often emerge from the environment — from distribution shift, spurious correlations, or subtle model degradation that nobody notices until it's too late. As Vyacheslav Zholudev of Sumsub puts it, at scale it's "very easy to lose the moment when your model starts to degrade."
And here's the insidious part: a model can be technically correct while practically wrong. Matt Pipke of physIQ describes models that "fit whatever they can fit"—latching onto spurious correlations, such as regional hay fever trends, rather than actual pathology. The cost function is satisfied. The patient is not.
Accountability Is Stewardship, Not Blame
The goal of accountability isn't to find someone to punish when things go wrong. It's to build systems where things are less likely to go wrong — and where recovery is swift when they do.
This requires three concrete pillars:
-
Clear ownership. Accountability can't float in the organizational ether. Someone must own the problem domain — not just the code. Mahyar Salek of Deeplook operationalizes this by embedding cytopathologists directly within the data science team, ensuring those who understand biological ground truth are responsible for model development. Patrick Leung of Earthshot Labs calls this the "danger zone" problem: a team of only software engineers and statisticians, with no domain expert to own the ecological reality, is an accountability disaster waiting to happen.
-
Full auditability. If a system fails, you must be able to trace why. This means versioning not just model weights but entire pipelines. It means data provenance — knowing exactly where data came from, who touched it, and which models were trained on it. David Sontag of Layer Health puts it simply: the system must be able to definitively state which exact model version produced a specific answer. Not approximately. Exactly.
-
Redress and recovery. Errors will happen. What matters is what you do next. Harro Stokman of Kepler Vision describes a feedback loop where false alarms are flagged, added to the training set, and used to update the model immediately. Amy Brown of Authenticx goes further: their interface lets users explicitly agree or disagree with AI predictions, using that human signal to continuously tune model behavior. The user isn't just a consumer of the AI — they're a check on it.
The Human-in-the-Loop Is Non-Negotiable (For Now)
There's a strong consensus among practitioners on where the red line currently sits: AI should assist, not autonomously decide, unless it has undergone the highest level of regulatory scrutiny.
Dirk Smeets of icometrix describes their tool as "a background service that empowers the radiologist"—it automates measurements and surfaces information, but the diagnosis remains with the clinician. Ranveer Chandra of Microsoft Research frames it similarly for agriculture: the vision is to augment a farmer's knowledge with data-driven insights, not to replace the farmer who understands the land.
This isn't just philosophical caution. It's risk architecture. When something goes wrong with a human decision, it's an isolated error. When something goes wrong with an autonomous AI system, it's a systematic failure — and the reputational, legal, and human costs scale accordingly.
For the rare cases where humans are removed from the loop entirely, the accountability burden simply shifts upstream. John Bertrand of Digital Diagnostics built an ethical framework into their system from inception, designing training data and clinical trials to account for diverse populations from day one — because once fully autonomous deployment begins, there is no safety net to catch what was missed.
Governance Is a Competitive Advantage
Here's the mindset shift that separates impactful AI teams from the rest: governance isn't red tape. It's quality assurance.
Matt Pipke of physIQ admits that the regulatory process is frustrating — but strict design controls made his team "better critics of their own work" and produced algorithms that far exceeded what a typical startup would build. Emi Gal of Ezra calls FDA clearance a "forcing function" that compels companies to validate performance across diverse populations from day one, rather than retrofitting safety later. Yiannis Kanellopoulos of Code4Thought advocates for independent last-mile analytics — audits performed outside the development team precisely because internal teams are optimized to solve business problems, not find their own blind spots.
The bottom line is that without accountability, there is no trust. Without trust, there is no adoption. Berk Birand of Fero Labs found that engineers in high-stakes manufacturing simply refuse to use a black box — because their jobs are on the line. Dean Freestone of Seer found the same in healthcare: without the signature of accountability, clinicians viewed AI as adding risk rather than reducing it.
The mandate is simple, if demanding: if you cannot trace your model's lineage and control its revisions, you shouldn't be deploying it.
- Heather
|
|
 |
|
Vision AI that bridges research and reality
— delivering where it matters
|
|
|
|
|
|
 |
Research: Foundation Models with Time Series
The Fourth Dimension: How Earth Observation AI is Mastering Time
For years, analyzing satellite imagery meant looking at static snapshots—treating the planet like a fixed map. But the Earth is dynamic, and the data is messy: clouds block views, orbits are irregular, and crops change weekly.
Three new papers illustrate a new capability in Earth Observation Foundation Models. The focus has moved from spatial features to spatio-temporal modeling—teaching AI to understand the "video" of our planet.
Here is how the latest research tackles the challenge of time:
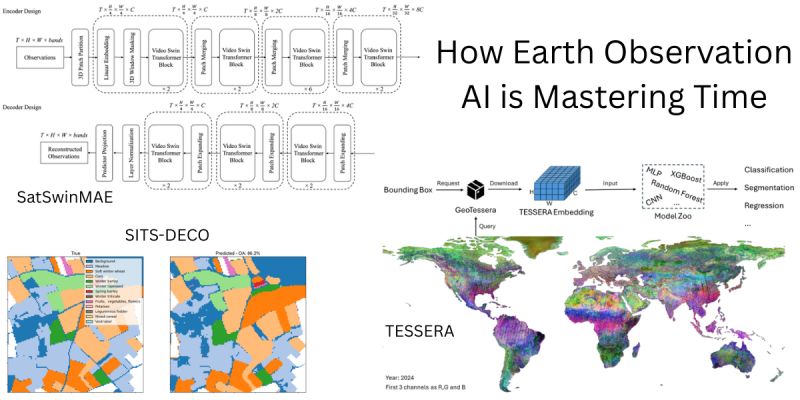
The GPT Moment for Satellites: Most EO models use encoders (like BERT), but Samuel Barrett and Docko Sow ask: what if we treated satellite data like language? They introduce SITS-DECO, a decoder-only generative model that processes satellite time series as unified token sequences. Inspired by LLMs, it uses symbolic prompting to perform multiple tasks (like crop classification) within a single architecture, without specific adaptation heads. Despite its simplicity, this generative approach outperforms much larger models, proving that dense temporal sequence modeling is the missing ingredient in current baselines.
Solving the Cloud & Orbit Chaos: Satellite data is notoriously irregular due to cloud cover and orbital gaps. Zhengpeng Feng et al. present TESSERA, a pixel-wise foundation model designed explicitly for this irregularity. Using Barlow Twins and sparse random temporal sampling, they enforce invariance to missing data. Their key innovation is a combination of global shuffling (to decorrelate spatial neighborhoods) and mix-based regulation to handle extreme sparsity. The team is releasing global, annual, 10m embeddings.
Adapting Video Tech to Earth: Yohei Nakayama et al. take a different route with SatSwinMAE, adapting state-of-the-art video processing for the Earth. They employ a Hierarchical 3D Masked Autoencoder with Video Swin Transformer blocks to capture multi-scale spatio-temporal dependencies. By adding U-Net-style skip connections to preserve scale-specific details, they achieve a large accuracy jump in land cover segmentation on the PhilEO Bench compared to existing geospatial foundation models.
The takeaway: The next generation of EO models won't just tell us what is on the ground. By mastering time series—whether through generative decoding (SITS-DECO), robust embedding (TESSERA), or 3D video attention (SatSwinMAE)—they will tell us how the ground is changing.
SITS-DECO: A Generative Decoder Is All You Need For Multitask Satellite Image Time Series Modelling
TESSERA: Temporal Embeddings of Surface Spectra for Earth Representation and Analysis
SatSwinMAE: Efficient Autoencoding for Multiscale Time-series Satellite Imagery
|
|
|
|
|
|
 |
Research: Foundation Model Robustness
Scanner-Induced Domain Shifts Undermine the Robustness of Pathology Foundation Models
We often evaluate pathology foundation models on benchmark AUCs and assume they are ready for the clinic. But what happens when you simply change the slide scanner? A new study reveals a critical vulnerability that accuracy metrics alone cannot show.
Real-world clinical deployment requires dealing with different hardware. If an AI model's embedding space fundamentally shifts just because a whole slide image (WSI) was digitized on a different machine, clinical reliability and generalization are at risk.
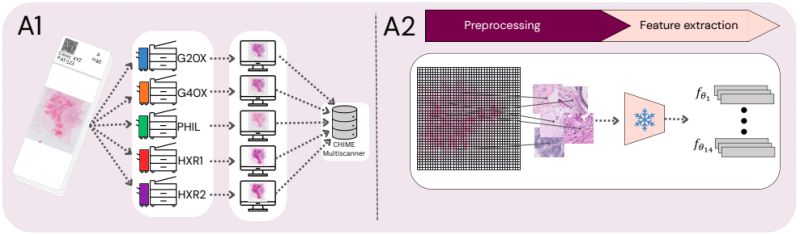
To investigate this, Erik Thiringer et al. systematically evaluated 14 pathology foundation models. By utilizing a multiscanner dataset where 384 breast cancer WSIs were digitized across five different devices, the team isolated scanner effects from underlying biological and laboratory confounders. In their assessment, they specifically focused on evaluating slide-level embeddings (via mean pooling) rather than just relying on patch-level analysis.
Here is what they found:
• While AUC often remains stable across scanners, this masks a deeper issue. The models encode pronounced scanner-specific variability that alters the embedding space and degrades the calibration of downstream predictions. The team confirmed this was not just spurious noise: slides from two physically different scanners of the 𝘴𝘢𝘮𝘦 model consistently clustered together, acting as a perfect positive control for genuine acquisition differences.
• Multimodal Vision-Language Models (like CONCH and CONCHv1.5) demonstrated the greatest scanner invariance. This might be due to their massive, heterogeneous training data (over a million PubMed image-caption pairs) regularizing the learned representations, or the multimodal alignment objective itself prioritizing semantic morphology over low-level scanner cues. 𝘏𝘰𝘸𝘦𝘷𝘦𝘳, despite winning on embedding consistency, they underperformed compared to other models on downstream predictive performance.
• Targeted training strategies help. The distilled model H0-mini consistently outperformed its massive teacher (H-Optimus-0) in cross-scanner embedding consistency. But while this robustness-focused distillation improved generalization without increasing model size, H0-mini still suffered from poor calibration.
𝗧𝗵𝗲 𝗧𝗮𝗸𝗲𝗮𝘄𝗮𝘆: None of the evaluated models provided reliable, out-of-the-box robustness against scanner-induced variability. The development of clinical-grade AI requires moving beyond accuracy-centric benchmarks and toward explicit evaluation of embedding stability and calibration under realistic hardware shifts.
|
|
|
|
|
|
 |
Research: Robustness
SCORPION: Addressing Scanner-Induced Variability in Histopathology
AI models in computational pathology often perform brilliantly in the lab, but what happens when a hospital uses a different digital scanner? The answer: predictions can shift, jeopardizing patient diagnosis and treatment planning.
Real-world adoption of digital pathology requires models to generalize across hardware. Previously, researchers tried to address this by evaluating models on unseen scanners during training. However, they lacked a direct way to test if a model produced consistent results for the 𝘦𝘹𝘢𝘤𝘵 𝘴𝘢𝘮𝘦 tissue when scanned across different devices.
Jeongun Ryu et al. just introduced a comprehensive solution to this problem. Here is how they are setting a new standard for reliability testing:
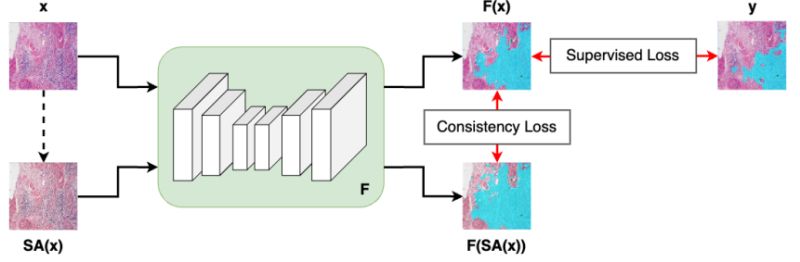
• 𝗧𝗵𝗲 𝗦𝗖𝗢𝗥𝗣𝗜𝗢𝗡 𝗗𝗮𝘁𝗮𝘀𝗲𝘁 (available for download): To strictly isolate hardware effects from biological differences, the team created a dataset of 480 tissue samples, each scanned across 5 different devices. This yields 2,400 spatially aligned patches, allowing researchers to rigorously evaluate model consistency while perfectly controlling for differences in tissue composition. Crucially, the authors are releasing this paired dataset to provide the research community with a standardized resource for reliability testing.
• 𝗧𝗵𝗲 𝗦𝗶𝗺𝗖𝗼𝗻𝘀 𝗙𝗿𝗮𝗺𝗲𝘄𝗼𝗿𝗸 & 𝗖𝗼𝗻𝘀𝗶𝘀𝘁𝗲𝗻𝗰𝘆 𝗟𝗼𝘀𝘀: To actively fix the variability, they propose SimCons, a framework that combines augmentation-based domain generalization techniques with a dedicated "consistency loss." By leveraging the spatially aligned patches of the exact same tissue from different scanners, this consistency loss explicitly penalizes the model if its outputs change depending on the scanning device. It effectively forces the model to ignore scanner-induced visual noise and anchor its predictions entirely on the underlying biology.
The results? SimCons successfully improves model consistency on varying scanners without compromising task-specific accuracy. By releasing the SCORPION dataset and proposing SimCons, the authors provide the research community with a crucial toolkit for building truly clinical-grade AI.
|
|
|
|
|
|
 |
Research: Weak Supervision
Making AI Clinical-Grade: Solving the Aggregation Problem
A single pathology slide contains billions of pixels. You cannot simply feed a whole slide image into a neural network; the file is too massive. Instead, we chop it into thousands of tiny tiles.
But a diagnosis is not a tile—it is the sum of the parts. This is where Multiple Instance Learning (MIL) comes in. MIL is the algorithmic brain that aggregates thousands of patch-level clues into a single patient-level answer.
While standard MIL works well in controlled experiments, it often breaks down in the messy reality of clinical deployment. Three new papers propose architectural shifts to handle the specific edge cases of real-world pathology: imbalance, forgetting, and efficiency.
Here is how the latest research upgrades the MIL framework:
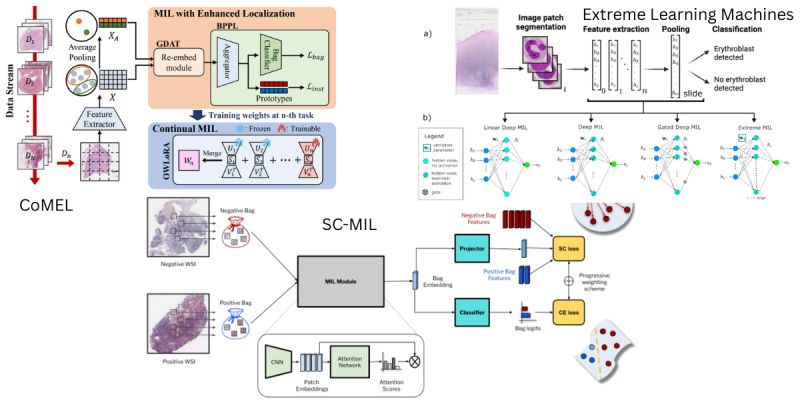
1) Real-world datasets are rarely balanced. Dinkar Juyal et al. (SC-MIL) address the double imbalance in pathology: rare disease classes occur infrequently, and even within a positive slide, tumor regions may constitute only a tiny fraction of the tissue. They integrate Supervised Contrastive Learning into the MIL framework, decoupling feature learning from classification. By forcing the model to learn robust decision boundaries before classification, they achieve significant gains, particularly in out-of-distribution settings.
2) Clinical AI is not static; it must learn new tasks over time without erasing old knowledge (catastrophic forgetting). Byung Hyun Lee et al. (CoMEL) argue that previous continual learning methods focused on global labels but failed to maintain localization accuracy (finding the cancer). Their framework allows models to adapt to new tasks while preserving spatial awareness. This approach outperformed prior methods in localization accuracy during sequential learning.
3) Does every task require massive compute? Rajiv Krishnakumar et al. challenge the reliance on heavy backpropagation. They combine attention-based MIL with Extreme Learning Machines—networks that use randomized, fixed weights for part of the architecture. Applied to the detection of circulating rare cells, this reduced the number of trained parameters by a factor of 5 while maintaining AUC within 1.5% of full deep learning models, proving that lightweight architectures can be robust enough for precision screening.
The takeaway: The path to deployment isn't just about larger foundation models; it's about specialized MIL architectures that can handle the scarcity, fluidity, and hardware constraints of the clinic.
SC-MIL: Supervised Contrastive Multiple Instance Learning for Imbalanced Classification in Pathology
Continual Multiple Instance Learning with Enhanced Localization for Histopathological Whole Slide Image Analysis
Extreme Learning Machines for Attention-based Multiple Instance Learning in Whole-Slide Image Classification |
|
|
|
|
|
|
Enjoy this newsletter? Here are more things you might find helpful:
Pixel Clarity Call - A free 30-minute conversation to cut through the noise and see where your vision AI project really stands. We’ll pinpoint vulnerabilities, clarify your biggest challenges, and decide if an assessment or diagnostic could save you time, money, and credibility.
Book now |
|
|
|
Did someone forward this email to you, and you want to sign up for more? Subscribe to future emails
This email was sent to _t.e.s.t_@example.com. Want to change to a different address? Update subscription
Want to get off this list? Unsubscribe
My postal address: Pixel Scientia Labs, LLC, PO Box 98412, Raleigh, NC 27624, United States |
|
|
|
|